Such models promise to transform biological research by providing a framework to (1) systematically interrogate and experimentally 
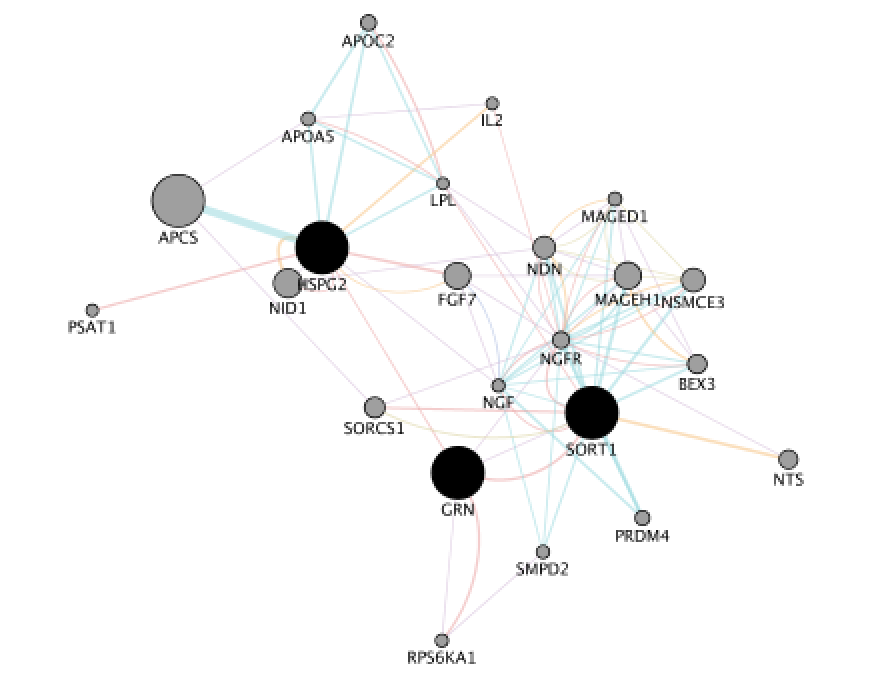
A variety of modeling environments haveīeen developed to simulate biochemical reactions and gene transcription kinetics ( Endy and Brent 2001), cellular physiology ( Loew and Schaff 2001), and metabolic control ( Mendes 1997). Interaction network for Halobacterium, and an interface to detailed stochastic/kinetic gene regulatory models.Ĭomputer-aided models of biological networks are a cornerstone of systems biology. SeveralĬase studies of Cytoscape plug-ins are surveyed, including a search for interaction pathways correlating with changes in geneĮxpression, a study of protein complexes involved in cellular recovery to DNA damage, inference of a combined physical/functional 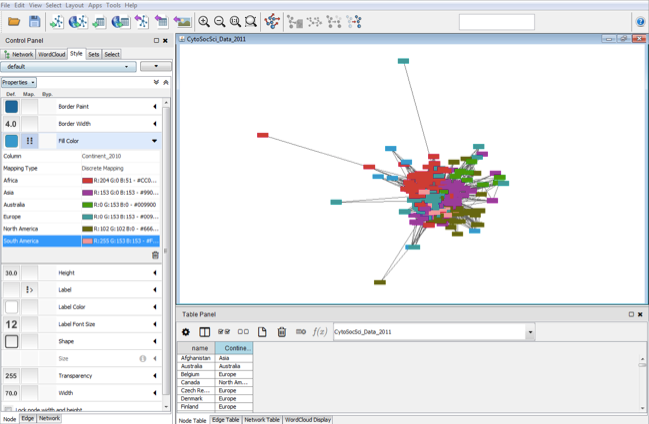
The Core is extensible throughĪ straightforward plug-in architecture, allowing rapid development of additional computational analyses and features. Cytoscape's software Core providesīasic functionality to layout and query the network to visually integrate the network with expression profiles, phenotypes,Īnd other molecular states and to link the network to databases of functional annotations. Although applicable to any system of molecular componentsĪnd interactions, Cytoscape is most powerful when used in conjunction with large databases of protein-protein, protein-DNA,Īnd genetic interactions that are increasingly available for humans and model organisms. The software architecture is based on the WSRF standard.Cytoscape is an open source software project for integrating biomolecular interaction networks with high-throughput expressionĭata and other molecular states into a unified conceptual framework. 
Lower-level computation will be performed through MPIBLAST. The Grid Service under development will analyse requests based on the number of sequences, splitting them accordingly to the available resources. These are the infrastructures that could reach the largest number of resources and the best load balancing for data access. Many efforts have been done in the literature concerning the speeding up of Blast searches, but few of them deal with the use of large heterogeneous production Grid Infrastructures. A prototype has been conceived for the particular problem of speeding up the Blast searches to obtain fast results for large datasets. In order to be able to offer the possibility of an enhanced computational capacity within this bioinformatics application, a Grid component is being developed. The power and analytical potential of both annotation and function data-mining is somehow restricted to the computational power behind each particular installation. Typical B2G users are middle size genomics labs carrying out sequencing, ETS and microarray projects, handling datasets up to several thousand sequences. The application has been developed with the aim of affering an easy-to-use tool for functional genomics research. Blast2GO (B2G) is a bioinformatics tool for Gene Ontology-based DNA or protein sequence annotation and function-based data mining. The vast amount in complexity of data generated in Genomic Research implies that new dedicated and powerful computational tools need to be developed to meet their analysis requirements. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
